Alignment Representation (Graph)¶
- Learning Objective
This tutorial introduces you to the graph data structures that can be used to represent an alignment in SeqAn. You will learn basic techniques to create and modify such data structures and how to access certain information from these data structures.
- Difficulty
Basic
- Duration
15 min
- Prerequisites
Another very useful representation of alignments is given by the Alignment Graph. It is a graph in which each vertex corresponds to a sequence segment, and each edge indicates an ungapped alignment between the connected vertices, or more precisely between the sequences stored in those vertices. Here is an example of such a graph:
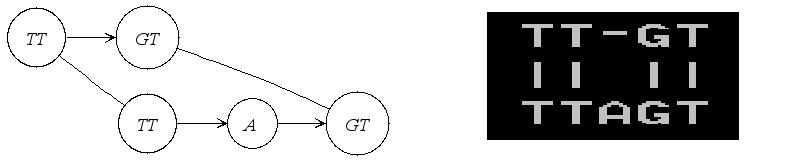
In the following we will actually construct this example step by step.
First we include the iostream header from the STL and the <seqan/align.h> header to include all necessary functions and data structures we want to use.
We use the namespace seqan and write the main function with an empty body.
#include <iostream>
#include <seqan/align.h>
using namespace seqan2;
int main()
{
At the begin of the function we define our types we want to use later on.
We define TSequence as the type of our input strings.
Since we work with a Dna alphabet we define TSequence as a String over a Dna alphabet.
For the AlignmentGraph we need two StringSets.
The TStringSet is used to actually store the input sequences and the TDepStringSet is internally used by the AlignmentGraph.
That is the AlignmentGraph does not copy the sources into its data structure but rather stores a reference to each of the given input strings as it does not modify the input sequences.
The Dependent StringSet facilitates this behavior.
In the end we define the actual AlignmentGraph type.
typedef String<Dna> TSequence;
typedef StringSet<TSequence> TStringSet;
typedef StringSet<TSequence, Dependent<> > TDepStringSet;
typedef Graph<Alignment<TDepStringSet> > TAlignGraph;
typedef typename VertexDescriptor<TAlignGraph>::Type TVertexDescriptor;
We first create our two input sequences TTGT and TTAGT append them to the StringSet strings using the appendValue function and pass the initialized strings object as a parameter to the constructor of the AlignmentGraph alignG.
TSequence seq1 = "TTGT";
TSequence seq2 = "TTAGT";
TStringSet strings;
appendValue(strings, seq1);
appendValue(strings, seq2);
TAlignGraph alignG(strings);
std::cout << alignG << std::endl;
Before adding vertices to the graph align prints the empty adjacency and edge
list.
Adjacency list:
Edge list:
Before we construct the alignment we print the unmodified AlignmentGraph.
Then we add some alignment information to the graph.
In order to add an ungapped alignment segment we have to add an edge connecting two vertices of different input sequences.
To do so we can use the function addEdge and specify the two vertices that should be connected.
Since we do not have any vertices yet, we create them on the fly using the function addVertex().
The function addVertex gets as second parameter the id which points to the the correct input sequence within the strings object.
We can use the function positionToId() to receive the id that corresponds to a certain position within the underlying Dependent StringSet of the AlignmentGraph.
We can access the Dependent StringSet using the function stringSet(). The third parameter of addVertex specifies the begin position of the segment within the respective input sequence and the fourth parameter specifies its length. Now, we add an edge between the two vertices of each input sequence which covers the first two positions. In the next step we have to add a gap. We can do this simply by just adding a vertex that covers the inserted string. Finally we have to add the second edge to represent the last ungapped sequence and then we print the constructed alignment.
Note that we use findVertex() to find the the last two inserted vertices. The syntax is the same as addVertex(), but omits the length parameter.
TVertexDescriptor u,v;
// TT
u = addVertex(alignG, positionToId(stringSet(alignG), 0), 0, 2);
v = addVertex(alignG, positionToId(stringSet(alignG), 1), 0, 2);
addEdge(alignG, u, v);
// A
addVertex(alignG, positionToId(stringSet(alignG), 1), 2, 1);
// GT
addVertex(alignG, positionToId(stringSet(alignG), 0), 2, 2);
addVertex(alignG, positionToId(stringSet(alignG), 1), 3, 2);
u = findVertex(alignG, positionToId(stringSet(alignG), 0), 2);
v = findVertex(alignG, positionToId(stringSet(alignG), 1), 3);
addEdge(alignG, u, v);
std::cout << alignG << std::endl;
return 0;
}
Now align prints the desired alignment.
Alignment matrix:
0 .
TT-GT
|| ||
TTAGT
The general usage of graphs is explained in the Graphs tutorial.
Assignment 1¶
- Type
Review
- Objective
Construct a multiple sequence alignment using the Alignment Graph data structure. Use the three sequences
GARFIELDTHECAT,GARFIELDTHEBIGCATandTHEBIGCATand align them such that you obtain the maximal number of matches.- Hints
TSequenceshould beString<char>instead ofString<Dna>.The function findVertex returns the vertex of an AlignmentGraph that covers the given position in the given sequence.
- Solution
#include <iostream> #include <seqan/align.h> using namespace seqan2; int main() { // Define the types we need. typedef String<char> TSequence; typedef StringSet<TSequence> TStringSet; typedef StringSet<TSequence, Dependent<> > TDepStringSet; typedef Graph<Alignment<TDepStringSet> > TAlignGraph; typedef typename VertexDescriptor<TAlignGraph>::Type TVertexDescriptor; // Initializing the sequences and the string set. TSequence seq1 = "GARFIELDTHECAT"; TSequence seq2 = "GARFIELDTHEBIGCAT"; TSequence seq3 = "THEBIGCAT"; TStringSet strings; appendValue(strings, seq1); appendValue(strings, seq2); appendValue(strings, seq3); // Load the string set into the Alignment Graph. TAlignGraph alignG(strings); TVertexDescriptor u,v; // Add two vertices covering "GARFIELD" in the first and the second sequence and connect them with an edge. u = addVertex(alignG, positionToId(stringSet(alignG), 0), 0, 8); v = addVertex(alignG, positionToId(stringSet(alignG), 1), 0, 8); addEdge(alignG, u, v); // Add two vertices covering "THE" in the first and the second sequence and connect them with an edge. u = addVertex(alignG, positionToId(stringSet(alignG), 0), 8, 3); v = addVertex(alignG, positionToId(stringSet(alignG), 1), 8, 3); addEdge(alignG, u, v); // Find the vertex covering "THE" in the first sequence and add the vertex covering "THE" in the third sequence and connect them with an edge. u = findVertex(alignG, positionToId(stringSet(alignG), 0), 8); v = addVertex(alignG, positionToId(stringSet(alignG), 2), 0, 3); addEdge(alignG, u, v); // Find the vertices covering "THE" in the second and the third sequence and connect them with an edge. u = findVertex(alignG, positionToId(stringSet(alignG), 1), 8); v = findVertex(alignG, positionToId(stringSet(alignG), 2), 0); addEdge(alignG, u, v); // Add two vertices covering "FAT" in the second and the third sequence and connect them with an edge. u = addVertex(alignG, positionToId(stringSet(alignG), 1), 11, 3); v = addVertex(alignG, positionToId(stringSet(alignG), 2), 3, 3); addEdge(alignG, u, v); // Add two vertices covering "CAT" in the first and the second sequence and connect them with an edge. u = addVertex(alignG, positionToId(stringSet(alignG), 0), 11, 3); v = addVertex(alignG, positionToId(stringSet(alignG), 1), 14, 3); addEdge(alignG, u, v); // Find the vertex covering "CAT" in the first sequence and add the vertex covering "CAT" in the third sequence and connect them with an edge. u = findVertex(alignG, positionToId(stringSet(alignG), 0), 11); v = addVertex(alignG, positionToId(stringSet(alignG), 2), 6, 3); addEdge(alignG, u, v); // Find the vertices covering "CAT" in the second and the third sequence and connect them with an edge. u = findVertex(alignG, positionToId(stringSet(alignG), 1), 14); v = findVertex(alignG, positionToId(stringSet(alignG), 2), 6); addEdge(alignG, u, v); std::cout << alignG << std::endl; return 0; }
Alignment matrix: 0 . : . GARFIELDTHE---CAT ||||||||||| ||| GARFIELDTHEBIGCAT ||||||||| --------THEBIGCAT